Reproducing McBride et al. (2023)
Overview
This guide documents a partitioned SARS-CoV-2 FLC analysis in white-tail deer, following McBride et al. (2023). The workflow uses episodic to run a Fixed Local Clock (FLC) analysis with multiple partitions corresponding to different genomic regions. The analysis focuses on comparing evolutionary rates across partitions and identifying accelerated evolution associated with host adaptation in deer.
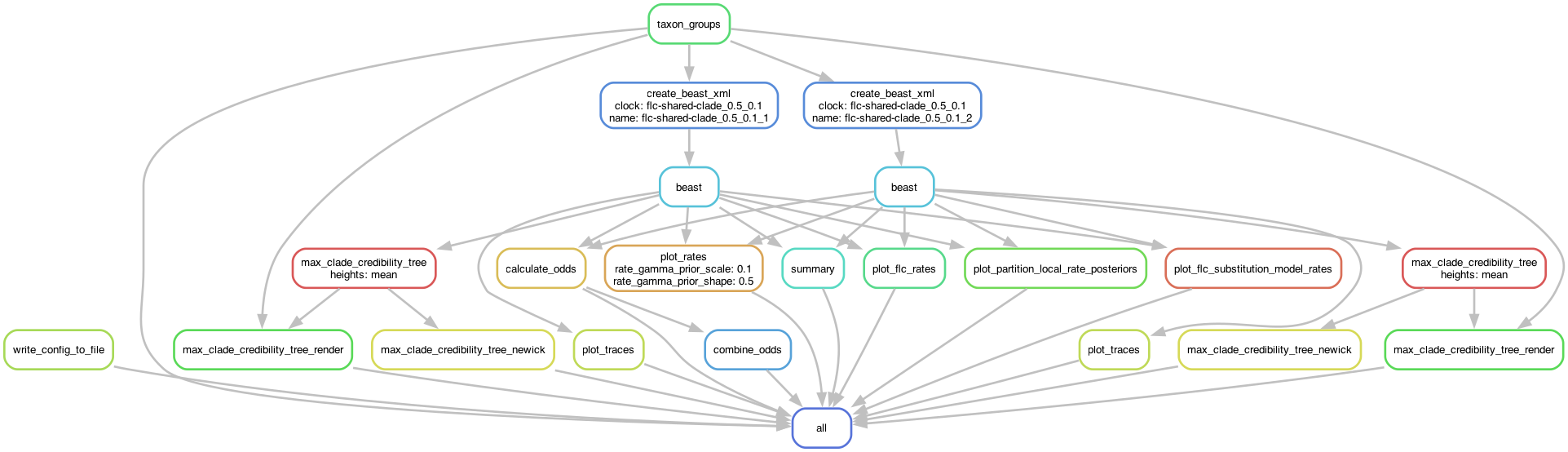
This analysis combine the previous guides on partitioning and FLC models to demonstrate how to set up a complex workflow with multiple partitions and a local-clock model.
A local clock root to tip regression of the dataset is available via the Clockor2 webapp. Root-to-tip regression is a useful exploratory tool for assessing temporal signal and clock-like behavior in the data, clockor2 allows you to interactively explore and fit local clock models to the data, which can inform the design of the full BEAST analysis.
Analysis setup
We have labeled white-tailed deer transmission clusters in the fasta headers according to McBride et al. (2023). For example, NODE_0000489 is a cluster of three deer sequences. As in McBride et al. (2023), we use a clade model here to match the biological hypothesis of sustained elevated rates within deer-associated clades, however a stem model could also be used if the hypothesis was focused on the transition period.
We use partitions to compare rates across different genomic regions. The concatenated accessory partition (ORF3-ORF8) contains ORF3a, envelope (E), membrane (M), ORF6, ORF7a, ORF7b, and ORF8. Episodic can extract these partitions from the original fasta file using the extract_sarscov2_partitions script, which uses SARS-CoV-2 partition definitions to generate one FASTA file per partition.
episodic script run \
extract_sarscov2_partitions \
--env python -- \
tests/data/McBride2023.fasta \
--concat-ORF3-E-M-ORF8 \
-o deer_partitions
Configure and Execute workflow
The run command below specifies the input partitions, local clock groups, and other key parameters. We run a single FLC model here, but you could easily add comparators by repeating the --clock option.
episodic run \
--output-dir McBride2023 \
--beast-chain-length 50000000 \
--clock flc-shared-clade \
--foreground-label Deer \
--background-label Human \
-a deer_partitions/N.fasta \
-a deer_partitions/ORF1a.fasta \
-a deer_partitions/ORF1b.fasta \
-a deer_partitions/S.fasta \
-a deer_partitions/ORF3-ORF8.fasta \
-g NODE_0000489 \
-g NODE_0000444 \
-g NODE_0000451 \
-g NODE_0000243 \
-g NODE_0000214 \
-g NODE_0000191 \
-g NODE_0000267 \
-g NODE_0000361 \
-g NODE_0000100 \
-g NODE_0000173 \
-g NODE_0000169 \
-g NODE_0000155 \
-g NODE_0000157 \
-g NODE_0000163
Key outputs
The posterior distributions of evolutionary rates (substitutions per site per year) for five partitions of the SARS-CoV-2 genome (ORF1a, ORF1b, ORF3–ORF8 plus envelope (E) and membrane (M), spike (S), and nucleocapsid (N)) are presented for human (pink) and WTD (blue) for the delta variant. Here we can see that the deer-associated clade has elevated rates across all partitions, which is consistent with the hypothesis of host adaptation driving accelerated evolution in deer.
