Partitioned H1N1 strict-clock analysis
Overview
This guide shows how to run a partitioned H1N1 HA analysis with episodic using a strict molecular clock and no FLC foreground groups.
Reference
This example follows the BEAST partitioning workflow pattern:
- https://beast.community/constructing_models#importing-partitions-from-multiple-files
Inputs
Primary input alignment:
tests/data/H1N1_HA.nex
Taxon labels encode sampling time using @ and may include uncertainty (for example, hSCar@1918/1).
The workflow parses this format using:
date_delimiter: '@'date_index: -1
Convert partitions to FASTA
The NEXUS file contains partition definitions (charset HA1, charset HA2). Extract one FASTA per partition:
episodic script run extract_partitions_from_nexus --env python -- tests/data/H1N1_HA.nex --output-dir tests/data
Expected files:
tests/data/HA1.fastatests/data/HA2.fasta
Analysis setup
- Model class: strict clock (
strict) - Partitions: HA1 and HA2
- Independent chains: duplicate BEAST runs
- Outputs: posterior summaries, trace diagnostics, MCC trees
Run
Validate configuration:
episodic run -a tests/data/HA1.fasta -a tests/data/HA2.fasta -o H1N1 -cl 1000000 -s 1000 --clock strict --dry
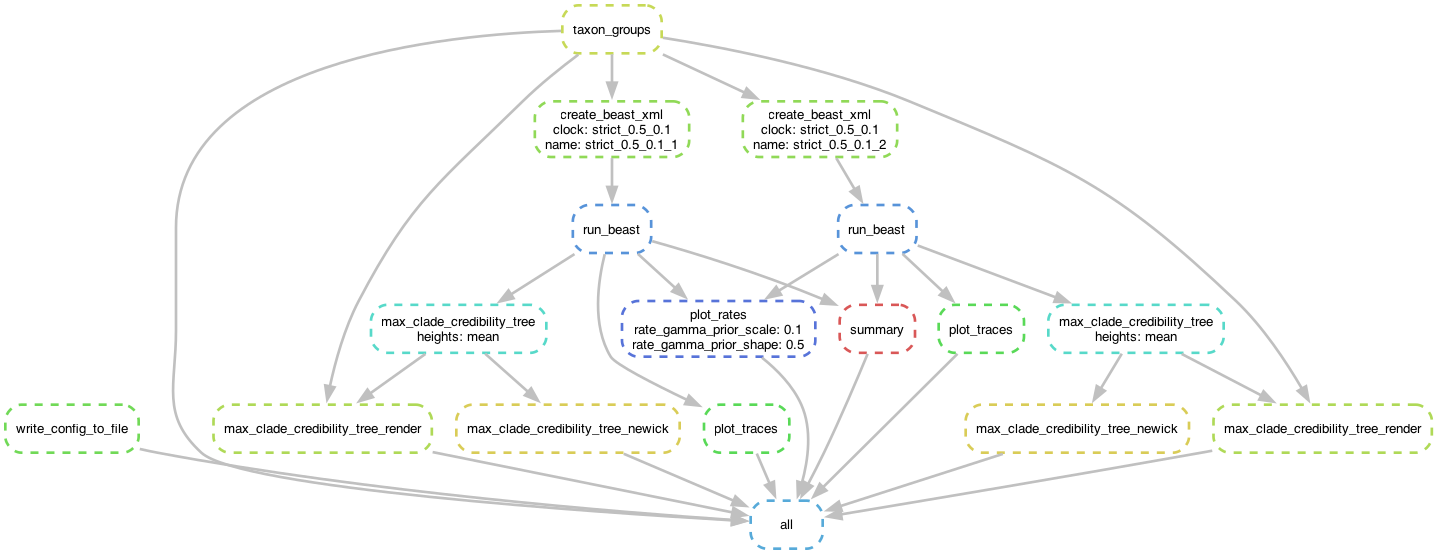
Generate workflow DAG:
episodic run -a tests/data/HA1.fasta -a tests/data/HA2.fasta -o H1N1 -cl 1000000 -s 1000 --clock strict --dag H1N1_dag.pdf
Run full analysis:
episodic run -a tests/data/HA1.fasta -a tests/data/HA2.fasta -o H1N1 -cl 1000000 -s 1000 --clock strict
Key outputs
For complete file naming patterns, see Workflow Outputs.
Trace comparison example across duplicate runs:
H1N1/clocks/strict_0.5_0.1/strict_0.5_0.1_1/strict_0.5_0.1_1_trace_plots/age(root).pngH1N1/clocks/strict_0.5_0.1/strict_0.5_0.1_2/strict_0.5_0.1_2_trace_plots/age(root).png
_1.png)
_2.png)
Compare the rates of the two partitions:
